kwcoco.metrics package¶
Submodules¶
- kwcoco.metrics.assignment module
- kwcoco.metrics.clf_report module
- kwcoco.metrics.confusion_measures module
- kwcoco.metrics.confusion_vectors module
- kwcoco.metrics.detect_metrics module
DetectionMetricsDetectionMetrics.clear()DetectionMetrics.enrich_confusion_vectors()DetectionMetrics.from_coco()DetectionMetrics._register_imagename()DetectionMetrics.add_predictions()DetectionMetrics.add_truth()DetectionMetrics.true_detections()DetectionMetrics.pred_detections()DetectionMetrics.classesDetectionMetrics.confusion_vectors()DetectionMetrics.score_kwant()DetectionMetrics.score_kwcoco()DetectionMetrics.score_voc()DetectionMetrics._to_coco()DetectionMetrics.score_pycocotools()DetectionMetrics.score_coco()DetectionMetrics.demo()DetectionMetrics.summarize()
_demo_construct_probs()pycocotools_confusion_vectors()eval_detections_cli()_summarize()pct_summarize2()
- kwcoco.metrics.drawing module
- kwcoco.metrics.functional module
- kwcoco.metrics.sklearn_alts module
- kwcoco.metrics.util module
- kwcoco.metrics.voc_metrics module
Module contents¶
mkinit kwcoco.metrics -w –relative
- class kwcoco.metrics.BinaryConfusionVectors(data, cx=None, classes=None)[source]¶
Bases:
NiceReprStores information about a binary classification problem. This is always with respect to a specific class, which is given by cx and classes.
- The data DataFrameArray must contain
is_true - if the row is an instance of class classes[cx] pred_score - the predicted probability of class classes[cx], and weight - sample weight of the example
Example
>>> from kwcoco.metrics.confusion_vectors import * # NOQA >>> self = BinaryConfusionVectors.demo(n=10) >>> print('self = {!r}'.format(self)) >>> print('measures = {}'.format(ub.urepr(self.measures())))
>>> self = BinaryConfusionVectors.demo(n=0) >>> print('measures = {}'.format(ub.urepr(self.measures())))
>>> self = BinaryConfusionVectors.demo(n=1) >>> print('measures = {}'.format(ub.urepr(self.measures())))
>>> self = BinaryConfusionVectors.demo(n=2) >>> print('measures = {}'.format(ub.urepr(self.measures())))
- classmethod demo(n=10, p_true=0.5, p_error=0.2, p_miss=0.0, rng=None)[source]¶
Create random data for tests
- Parameters:
n (int) – number of rows
p_true (float) – fraction of real positive cases
p_error (float) – probability of making a recoverable mistake
p_miss (float) – probability of making a unrecoverable mistake
rng (int | RandomState | None) – random seed / state
- Returns:
BinaryConfusionVectors
Example
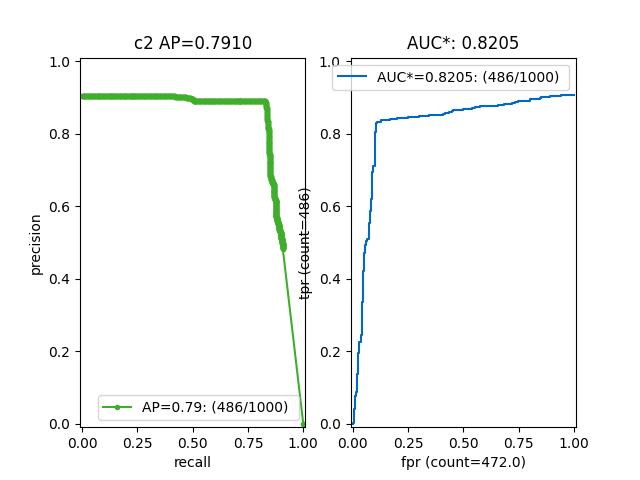
>>> from kwcoco.metrics.confusion_vectors import * # NOQA >>> cfsn = BinaryConfusionVectors.demo(n=1000, p_error=0.1, p_miss=0.1) >>> measures = cfsn.measures() >>> print('measures = {}'.format(ub.urepr(measures, nl=1))) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, pnum=(1, 2, 1)) >>> measures.draw('pr') >>> kwplot.figure(fnum=1, pnum=(1, 2, 2)) >>> measures.draw('roc')
- property catname¶
- measures(stabalize_thresh=7, fp_cutoff=None, monotonic_ppv=True, ap_method='pycocotools')[source]¶
Get statistics (F1, G1, MCC) versus thresholds
- Parameters:
stabalize_thresh (int, default=7) – if fewer than this many data points inserts dummy stabalization data so curves can still be drawn.
fp_cutoff (int | None) – maximum number of false positives in the truncated roc curves. The default of
Noneis equivalent tofloat('inf')monotonic_ppv (bool) – if True ensures that precision is always increasing as recall decreases. This is done in pycocotools scoring, but I’m not sure its a good idea.
Example
>>> from kwcoco.metrics.confusion_vectors import * # NOQA >>> self = BinaryConfusionVectors.demo(n=0) >>> print('measures = {}'.format(ub.urepr(self.measures()))) >>> self = BinaryConfusionVectors.demo(n=1, p_true=0.5, p_error=0.5) >>> print('measures = {}'.format(ub.urepr(self.measures()))) >>> self = BinaryConfusionVectors.demo(n=3, p_true=0.5, p_error=0.5) >>> print('measures = {}'.format(ub.urepr(self.measures())))
>>> self = BinaryConfusionVectors.demo(n=100, p_true=0.5, p_error=0.5, p_miss=0.3) >>> print('measures = {}'.format(ub.urepr(self.measures()))) >>> print('measures = {}'.format(ub.urepr(ub.odict(self.measures()))))
References
https://en.wikipedia.org/wiki/Confusion_matrix https://en.wikipedia.org/wiki/Precision_and_recall https://en.wikipedia.org/wiki/Matthews_correlation_coefficient
- _binary_clf_curves(stabalize_thresh=7, fp_cutoff=None)[source]¶
Compute TP, FP, TN, and FN counts for this binary confusion vector.
Code common to ROC, PR, and threshold measures, computes the elements of the binary confusion matrix at all relevant operating point thresholds.
- Parameters:
stabalize_thresh (int) – if fewer than this many data points insert stabalization data.
fp_cutoff (int | None) – maximum number of false positives
Example
>>> from kwcoco.metrics.confusion_vectors import * # NOQA >>> self = BinaryConfusionVectors.demo(n=1, p_true=0.5, p_error=0.5) >>> self._binary_clf_curves()
>>> self = BinaryConfusionVectors.demo(n=0, p_true=0.5, p_error=0.5) >>> self._binary_clf_curves()
>>> self = BinaryConfusionVectors.demo(n=100, p_true=0.5, p_error=0.5) >>> self._binary_clf_curves()
- _3dplot()[source]¶
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(module:pandas) >>> from kwcoco.metrics.confusion_vectors import * # NOQA >>> from kwcoco.metrics.detect_metrics import DetectionMetrics >>> dmet = DetectionMetrics.demo( >>> n_fp=(0, 1), n_fn=(0, 2), nimgs=256, nboxes=(0, 10), >>> bbox_noise=10, >>> classes=1) >>> cfsn_vecs = dmet.confusion_vectors() >>> self = bin_cfsn = cfsn_vecs.binarize_classless() >>> #dmet.summarize(plot=True) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=3) >>> self._3dplot()
- class kwcoco.metrics.ConfusionVectors(data, classes, probs=None)[source]¶
Bases:
NiceReprStores information used to construct a confusion matrix. This includes corresponding vectors of predicted labels, true labels, sample weights, etc…
- Variables:
data (kwarray.DataFrameArray) – should at least have keys true, pred, weight
classes (Sequence | CategoryTree) – list of category names or category graph
probs (ndarray | None) – probabilities for each class
Example
>>> # xdoctest: IGNORE_WANT >>> # xdoctest: +REQUIRES(module:pandas) >>> from kwcoco.metrics import DetectionMetrics >>> dmet = DetectionMetrics.demo( >>> nimgs=10, nboxes=(0, 10), n_fp=(0, 1), classes=3) >>> cfsn_vecs = dmet.confusion_vectors() >>> print(cfsn_vecs.data._pandas()) pred true score weight iou txs pxs gid 0 2 2 10.0000 1.0000 1.0000 0 4 0 1 2 2 7.5025 1.0000 1.0000 1 3 0 2 1 1 5.0050 1.0000 1.0000 2 2 0 3 3 -1 2.5075 1.0000 -1.0000 -1 1 0 4 2 -1 0.0100 1.0000 -1.0000 -1 0 0 5 -1 2 0.0000 1.0000 -1.0000 3 -1 0 6 -1 2 0.0000 1.0000 -1.0000 4 -1 0 7 2 2 10.0000 1.0000 1.0000 0 5 1 8 2 2 8.0020 1.0000 1.0000 1 4 1 9 1 1 6.0040 1.0000 1.0000 2 3 1 .. ... ... ... ... ... ... ... ... 62 -1 2 0.0000 1.0000 -1.0000 7 -1 7 63 -1 3 0.0000 1.0000 -1.0000 8 -1 7 64 -1 1 0.0000 1.0000 -1.0000 9 -1 7 65 1 -1 10.0000 1.0000 -1.0000 -1 0 8 66 1 1 0.0100 1.0000 1.0000 0 1 8 67 3 -1 10.0000 1.0000 -1.0000 -1 3 9 68 2 2 6.6700 1.0000 1.0000 0 2 9 69 2 2 3.3400 1.0000 1.0000 1 1 9 70 3 -1 0.0100 1.0000 -1.0000 -1 0 9 71 -1 2 0.0000 1.0000 -1.0000 2 -1 9
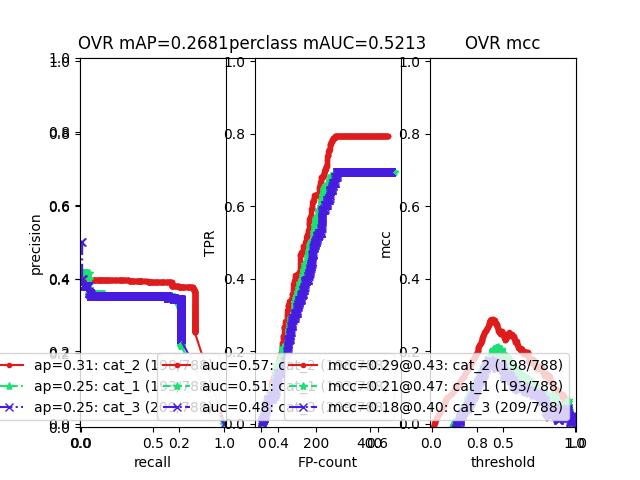
>>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:pandas) >>> import kwplot >>> kwplot.autompl() >>> from kwcoco.metrics.confusion_vectors import ConfusionVectors >>> cfsn_vecs = ConfusionVectors.demo( >>> nimgs=128, nboxes=(0, 10), n_fp=(0, 3), n_fn=(0, 3), classes=3) >>> cx_to_binvecs = cfsn_vecs.binarize_ovr() >>> measures = cx_to_binvecs.measures()['perclass'] >>> print('measures = {!r}'.format(measures)) measures = <PerClass_Measures({ 'cat_1': <Measures({'ap': 0.227, 'auc': 0.507, 'catname': cat_1, 'max_f1': f1=0.45@0.47, 'nsupport': 788.000})>, 'cat_2': <Measures({'ap': 0.288, 'auc': 0.572, 'catname': cat_2, 'max_f1': f1=0.51@0.43, 'nsupport': 788.000})>, 'cat_3': <Measures({'ap': 0.225, 'auc': 0.484, 'catname': cat_3, 'max_f1': f1=0.46@0.40, 'nsupport': 788.000})>, }) at 0x7facf77bdfd0> >>> kwplot.figure(fnum=1, doclf=True) >>> measures.draw(key='pr', fnum=1, pnum=(1, 3, 1)) >>> measures.draw(key='roc', fnum=1, pnum=(1, 3, 2)) >>> measures.draw(key='mcc', fnum=1, pnum=(1, 3, 3)) ...
- classmethod demo(**kw)[source]¶
- Parameters:
**kwargs – See
kwcoco.metrics.DetectionMetrics.demo()- Returns:
ConfusionVectors
Example
>>> from kwcoco.metrics.confusion_vectors import * # NOQA >>> cfsn_vecs = ConfusionVectors.demo() >>> print('cfsn_vecs = {!r}'.format(cfsn_vecs)) >>> cx_to_binvecs = cfsn_vecs.binarize_ovr() >>> print('cx_to_binvecs = {!r}'.format(cx_to_binvecs))
- classmethod from_arrays(true, pred=None, score=None, weight=None, probs=None, classes=None)[source]¶
Construct confusion vector data structure from component arrays
Example
>>> # xdoctest: +REQUIRES(module:pandas) >>> import kwarray >>> classes = ['person', 'vehicle', 'object'] >>> rng = kwarray.ensure_rng(0) >>> true = (rng.rand(10) * len(classes)).astype(int) >>> probs = rng.rand(len(true), len(classes)) >>> cfsn_vecs = ConfusionVectors.from_arrays(true=true, probs=probs, classes=classes) >>> cfsn_vecs.confusion_matrix() pred person vehicle object real person 0 0 0 vehicle 2 4 1 object 2 1 0
- confusion_matrix(compress=False)[source]¶
Builds a confusion matrix from the confusion vectors.
- Parameters:
compress (bool, default=False) – if True removes rows / columns with no entries
- Returns:
- cmthe labeled confusion matrix
- (Note: we should write a efficient replacement for
this use case. #remove_pandas)
- Return type:
pd.DataFrame
CommandLine
xdoctest -m kwcoco.metrics.confusion_vectors ConfusionVectors.confusion_matrix
Example
>>> # xdoctest: +REQUIRES(module:pandas) >>> from kwcoco.metrics import DetectionMetrics >>> dmet = DetectionMetrics.demo( >>> nimgs=10, nboxes=(0, 10), n_fp=(0, 1), n_fn=(0, 1), >>> classes=3, cls_noise=.2, newstyle=False) >>> cfsn_vecs = dmet.confusion_vectors() >>> cm = cfsn_vecs.confusion_matrix() ... >>> print(cm.to_string(float_format=lambda x: '%.2f' % x)) pred background cat_1 cat_2 cat_3 real background 0.00 1.00 2.00 3.00 cat_1 3.00 12.00 0.00 0.00 cat_2 3.00 0.00 14.00 0.00 cat_3 2.00 0.00 0.00 17.00
- binarize_classless(negative_classes=None)[source]¶
Creates a binary representation useful for measuring the performance of detectors. It is assumed that scores of “positive” classes should be high and “negative” clases should be low.
- Parameters:
negative_classes (List[str | int] | None) – list of negative class names or idxs, by default chooses any class with a true class index of -1. These classes should ideally have low scores.
- Returns:
BinaryConfusionVectors
Note
The “classlessness” of this depends on the compat=”all” argument being used when constructing confusion vectors, otherwise it becomes something like a macro-average because the class information was used in deciding which true and predicted boxes were allowed to match.
Example
>>> from kwcoco.metrics import DetectionMetrics >>> dmet = DetectionMetrics.demo( >>> nimgs=10, nboxes=(0, 10), n_fp=(0, 1), n_fn=(0, 1), classes=3) >>> cfsn_vecs = dmet.confusion_vectors() >>> class_idxs = list(dmet.classes.node_to_idx.values()) >>> binvecs = cfsn_vecs.binarize_classless()
- binarize_ovr(mode=1, keyby='name', ignore_classes={'ignore'}, approx=False)[source]¶
Transforms cfsn_vecs into one-vs-rest BinaryConfusionVectors for each category.
- Parameters:
mode (int, default=1) – 0 for heirarchy aware or 1 for voc like. MODE 0 IS PROBABLY BROKEN
keyby (int | str) – can be cx or name
ignore_classes (Set[str]) – category names to ignore
approx (bool, default=0) – if True try and approximate missing scores otherwise assume they are irrecoverable and use -inf
- Returns:
- which behaves like
Dict[int, BinaryConfusionVectors]: cx_to_binvecs
- Return type:
Example
>>> from kwcoco.metrics.confusion_vectors import * # NOQA >>> cfsn_vecs = ConfusionVectors.demo() >>> print('cfsn_vecs = {!r}'.format(cfsn_vecs)) >>> catname_to_binvecs = cfsn_vecs.binarize_ovr(keyby='name') >>> print('catname_to_binvecs = {!r}'.format(catname_to_binvecs))
cfsn_vecs.data.pandas() catname_to_binvecs.cx_to_binvecs[‘class_1’].data.pandas()
Note
- class kwcoco.metrics.DetectionMetrics(classes=None)[source]¶
Bases:
NiceReprObject that computes associations between detections and can convert them into sklearn-compatible representations for scoring.
- Variables:
gid_to_true_dets (Dict[int, kwimage.Detections]) – maps image ids to truth
gid_to_pred_dets (Dict[int, kwimage.Detections]) – maps image ids to predictions
classes (kwcoco.CategoryTree | None) – the categories to be scored, if unspecified attempts to determine these from the truth detections
Example
>>> # Demo how to use detection metrics directly given detections only >>> # (no kwcoco file required) >>> from kwcoco.metrics import detect_metrics >>> import kwimage >>> # Setup random true detections (these are just boxes and scores) >>> true_dets = kwimage.Detections.random(3) >>> # Peek at the simple internals of a detections object >>> print('true_dets.data = {}'.format(ub.urepr(true_dets.data, nl=1))) >>> # Create similar but different predictions >>> true_subset = true_dets.take([1, 2]).warp(kwimage.Affine.coerce({'scale': 1.1})) >>> false_positive = kwimage.Detections.random(3) >>> pred_dets = kwimage.Detections.concatenate([true_subset, false_positive]) >>> dmet = DetectionMetrics() >>> dmet.add_predictions(pred_dets, imgname='image1') >>> dmet.add_truth(true_dets, imgname='image1') >>> # Raw confusion vectors >>> cfsn_vecs = dmet.confusion_vectors() >>> print(cfsn_vecs.data.pandas().to_string()) >>> # Our scoring definition (derived from confusion vectors) >>> print(dmet.score_kwcoco()) >>> # VOC scoring >>> print(dmet.score_voc(bias=0)) >>> # Original pycocotools scoring >>> # xdoctest: +REQUIRES(module:pycocotools) >>> print(dmet.score_pycocotools())
Example
>>> dmet = DetectionMetrics.demo( >>> nimgs=100, nboxes=(0, 3), n_fp=(0, 1), classes=8, score_noise=0.9, hacked=False) >>> print(dmet.score_kwcoco(bias=0, compat='mutex', prioritize='iou')['mAP']) ... >>> # NOTE: IN GENERAL NETHARN AND VOC ARE NOT THE SAME >>> print(dmet.score_voc(bias=0)['mAP']) 0.8582... >>> #print(dmet.score_coco()['mAP'])
- enrich_confusion_vectors(cfsn_vecs)[source]¶
Adds annotation id information into confusion vectors computed via this detection metrics object.
TODO: should likely use this at the end of the function that builds the confusion vectors.
- classmethod from_coco(true_coco, pred_coco, gids=None, verbose=0)[source]¶
Create detection metrics from two coco files representing the truth and predictions.
- Parameters:
true_coco (kwcoco.CocoDataset) – coco dataset with ground truth
pred_coco (kwcoco.CocoDataset) – coco dataset with predictions
Example
>>> import kwcoco >>> from kwcoco.demo.perterb import perterb_coco >>> true_coco = kwcoco.CocoDataset.demo('shapes') >>> perterbkw = dict(box_noise=0.5, cls_noise=0.5, score_noise=0.5) >>> pred_coco = perterb_coco(true_coco, **perterbkw) >>> self = DetectionMetrics.from_coco(true_coco, pred_coco) >>> self.score_voc()
- add_predictions(pred_dets, imgname=None, gid=None)[source]¶
Register/Add predicted detections for an image
- Parameters:
pred_dets (kwimage.Detections) – predicted detections
imgname (str | None) – a unique string to identify the image
gid (int | None) – the integer image id if known
- add_truth(true_dets, imgname=None, gid=None)[source]¶
Register/Add groundtruth detections for an image
- Parameters:
true_dets (kwimage.Detections) – groundtruth
imgname (str | None) – a unique string to identify the image
gid (int | None) – the integer image id if known
- property classes¶
- confusion_vectors(iou_thresh=0.5, bias=0, gids=None, compat='mutex', prioritize='iou', ignore_classes='ignore', background_class=NoParam, verbose='auto', workers=0, track_probs='try', max_dets=None)[source]¶
Assigns predicted boxes to the true boxes so we can transform the detection problem into a classification problem for scoring.
- Parameters:
iou_thresh (float | List[float]) – bounding box overlap iou threshold required for assignment if a list, then return type is a dict. Defaults to 0.5
bias (float) – for computing bounding box overlap, either 1 or 0 Defaults to 0.
gids (List[int] | None) – which subset of images ids to compute confusion metrics on. If not specified all images are used. Defaults to None.
compat (str) – can be (‘ancestors’ | ‘mutex’ | ‘all’). determines which pred boxes are allowed to match which true boxes. If ‘mutex’, then pred boxes can only match true boxes of the same class. If ‘ancestors’, then pred boxes can match true boxes that match or have a coarser label. If ‘all’, then any pred can match any true, regardless of its category label. Defaults to all.
prioritize (str) – can be (‘iou’ | ‘class’ | ‘correct’) determines which box to assign to if mutiple true boxes overlap a predicted box. if prioritize is iou, then the true box with maximum iou (above iou_thresh) will be chosen. If prioritize is class, then it will prefer matching a compatible class above a higher iou. If prioritize is correct, then ancestors of the true class are preferred over descendents of the true class, over unreleated classes. Default to ‘iou’
ignore_classes (set | str) – class names indicating ignore regions. Default={‘ignore’}
background_class (str | NoParamType) – Name of the background class. If unspecified we try to determine it with heuristics. A value of None means there is no background class.
verbose (int | str) – verbosity flag. Default to ‘auto’. In auto mode, verbose=1 if len(gids) > 1000.
workers (int) – number of parallel assignment processes. Defaults to 0
track_probs (str) – can be ‘try’, ‘force’, or False. if truthy, we assume probabilities for multiple classes are available. default=’try’
- Returns:
ConfusionVectors | Dict[float, ConfusionVectors]
Example
>>> dmet = DetectionMetrics.demo(nimgs=30, classes=3, >>> nboxes=10, n_fp=3, box_noise=10, >>> with_probs=False) >>> iou_to_cfsn = dmet.confusion_vectors(iou_thresh=[0.3, 0.5, 0.9]) >>> for t, cfsn in iou_to_cfsn.items(): >>> print('t = {!r}'.format(t)) ... print(cfsn.binarize_ovr().measures()) ... print(cfsn.binarize_classless().measures())
- score_kwcoco(iou_thresh=0.5, bias=0, gids=None, compat='all', prioritize='iou')[source]¶
our scoring method
- score_voc(iou_thresh=0.5, bias=1, method='voc2012', gids=None, ignore_classes='ignore')[source]¶
score using voc method
Example
>>> dmet = DetectionMetrics.demo( >>> nimgs=100, nboxes=(0, 3), n_fp=(0, 1), classes=8, >>> score_noise=.5) >>> print(dmet.score_voc()['mAP']) 0.9399...
- _to_coco()[source]¶
Convert to a coco representation of truth and predictions
with inverse aid mappings
- score_pycocotools(with_evaler=False, with_confusion=False, verbose=0, iou_thresholds=None)[source]¶
score using ms-coco method
- Returns:
dictionary with pct info
- Return type:
Dict
Example
>>> # xdoctest: +REQUIRES(module:pycocotools) >>> from kwcoco.metrics.detect_metrics import * >>> dmet = DetectionMetrics.demo( >>> nimgs=10, nboxes=(0, 3), n_fn=(0, 1), n_fp=(0, 1), classes=8, with_probs=False) >>> pct_info = dmet.score_pycocotools(verbose=1, >>> with_evaler=True, >>> with_confusion=True, >>> iou_thresholds=[0.5, 0.9]) >>> evaler = pct_info['evaler'] >>> iou_to_cfsn_vecs = pct_info['iou_to_cfsn_vecs'] >>> for iou_thresh in iou_to_cfsn_vecs.keys(): >>> print('iou_thresh = {!r}'.format(iou_thresh)) >>> cfsn_vecs = iou_to_cfsn_vecs[iou_thresh] >>> ovr_measures = cfsn_vecs.binarize_ovr().measures() >>> print('ovr_measures = {}'.format(ub.urepr(ovr_measures, nl=1, precision=4)))
Note
by default pycocotools computes average precision as the literal average of computed precisions at 101 uniformly spaced recall thresholds.
pycocoutils seems to only allow predictions with the same category as the truth to match those truth objects. This should be the same as calling dmet.confusion_vectors with compat = mutex
pycocoutils does not take into account the fact that each box often has a score for each category.
pycocoutils will be incorrect if any annotation has an id of 0
a major difference in the way kwcoco scores versus pycocoutils is the calculation of AP. The assignment between truth and predicted detections produces similar enough results. Given our confusion vectors we use the scikit-learn definition of AP, whereas pycocoutils seems to compute precision and recall — more or less correctly — but then it resamples the precision at various specified recall thresholds (in the accumulate function, specifically how pr is resampled into the q array). This can lead to a large difference in reported scores.
pycocoutils also smooths out the precision such that it is monotonic decreasing, which might not be the best idea.
pycocotools area ranges are inclusive on both ends, that means the “small” and “medium” truth selections do overlap somewhat.
- score_coco(with_evaler=False, with_confusion=False, verbose=0, iou_thresholds=None)¶
score using ms-coco method
- Returns:
dictionary with pct info
- Return type:
Dict
Example
>>> # xdoctest: +REQUIRES(module:pycocotools) >>> from kwcoco.metrics.detect_metrics import * >>> dmet = DetectionMetrics.demo( >>> nimgs=10, nboxes=(0, 3), n_fn=(0, 1), n_fp=(0, 1), classes=8, with_probs=False) >>> pct_info = dmet.score_pycocotools(verbose=1, >>> with_evaler=True, >>> with_confusion=True, >>> iou_thresholds=[0.5, 0.9]) >>> evaler = pct_info['evaler'] >>> iou_to_cfsn_vecs = pct_info['iou_to_cfsn_vecs'] >>> for iou_thresh in iou_to_cfsn_vecs.keys(): >>> print('iou_thresh = {!r}'.format(iou_thresh)) >>> cfsn_vecs = iou_to_cfsn_vecs[iou_thresh] >>> ovr_measures = cfsn_vecs.binarize_ovr().measures() >>> print('ovr_measures = {}'.format(ub.urepr(ovr_measures, nl=1, precision=4)))
Note
by default pycocotools computes average precision as the literal average of computed precisions at 101 uniformly spaced recall thresholds.
pycocoutils seems to only allow predictions with the same category as the truth to match those truth objects. This should be the same as calling dmet.confusion_vectors with compat = mutex
pycocoutils does not take into account the fact that each box often has a score for each category.
pycocoutils will be incorrect if any annotation has an id of 0
a major difference in the way kwcoco scores versus pycocoutils is the calculation of AP. The assignment between truth and predicted detections produces similar enough results. Given our confusion vectors we use the scikit-learn definition of AP, whereas pycocoutils seems to compute precision and recall — more or less correctly — but then it resamples the precision at various specified recall thresholds (in the accumulate function, specifically how pr is resampled into the q array). This can lead to a large difference in reported scores.
pycocoutils also smooths out the precision such that it is monotonic decreasing, which might not be the best idea.
pycocotools area ranges are inclusive on both ends, that means the “small” and “medium” truth selections do overlap somewhat.
- classmethod demo(**kwargs)[source]¶
Creates random true boxes and predicted boxes that have some noisy offset from the truth.
- Kwargs:
- classes (int):
class list or the number of foreground classes. Defaults to 1.
nimgs (int): number of images in the coco datasts. Defaults to 1.
nboxes (int): boxes per image. Defaults to 1.
n_fp (int): number of false positives. Defaults to 0.
- n_fn (int):
number of false negatives. Defaults to 0.
- box_noise (float):
std of a normal distribution used to perterb both box location and box size. Defaults to 0.
- cls_noise (float):
probability that a class label will change. Must be within 0 and 1. Defaults to 0.
- anchors (ndarray):
used to create random boxes. Defaults to None.
- null_pred (bool):
if True, predicted classes are returned as null, which means only localization scoring is suitable. Defaults to 0.
- with_probs (bool):
if True, includes per-class probabilities with predictions Defaults to 1.
rng (int | None | RandomState): random seed / state
CommandLine
xdoctest -m kwcoco.metrics.detect_metrics DetectionMetrics.demo:2 --show
Example
>>> kwargs = {} >>> # Seed the RNG >>> kwargs['rng'] = 0 >>> # Size parameters determine how big the data is >>> kwargs['nimgs'] = 5 >>> kwargs['nboxes'] = 7 >>> kwargs['classes'] = 11 >>> # Noise parameters perterb predictions further from the truth >>> kwargs['n_fp'] = 3 >>> kwargs['box_noise'] = 0.1 >>> kwargs['cls_noise'] = 0.5 >>> dmet = DetectionMetrics.demo(**kwargs) >>> print('dmet.classes = {}'.format(dmet.classes)) dmet.classes = <CategoryTree(nNodes=12, maxDepth=3, maxBreadth=4...)> >>> # Can grab kwimage.Detection object for any image >>> print(dmet.true_detections(gid=0)) <Detections(4)> >>> print(dmet.pred_detections(gid=0)) <Detections(7)>
Example
>>> # Test case with null predicted categories >>> dmet = DetectionMetrics.demo(nimgs=30, null_pred=1, classes=3, >>> nboxes=10, n_fp=3, box_noise=0.1, >>> with_probs=False) >>> dmet.gid_to_pred_dets[0].data >>> dmet.gid_to_true_dets[0].data >>> cfsn_vecs = dmet.confusion_vectors() >>> binvecs_ovr = cfsn_vecs.binarize_ovr() >>> binvecs_per = cfsn_vecs.binarize_classless() >>> measures_per = binvecs_per.measures() >>> measures_ovr = binvecs_ovr.measures() >>> print('measures_per = {!r}'.format(measures_per)) >>> print('measures_ovr = {!r}'.format(measures_ovr)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> measures_ovr['perclass'].draw(key='pr', fnum=2)
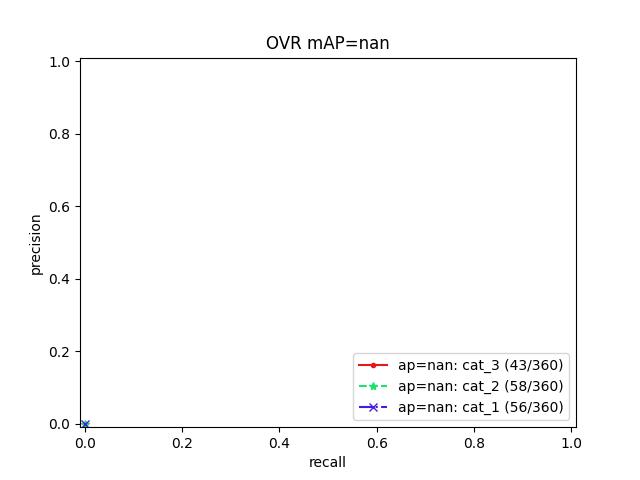
Example
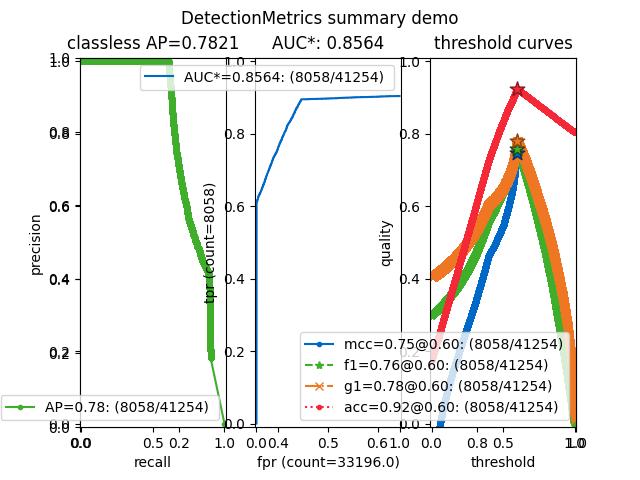
>>> from kwcoco.metrics.confusion_vectors import * # NOQA >>> from kwcoco.metrics.detect_metrics import DetectionMetrics >>> dmet = DetectionMetrics.demo( >>> n_fp=(0, 1), n_fn=(0, 1), nimgs=32, nboxes=(0, 16), >>> classes=3, rng=0, newstyle=1, box_noise=0.5, cls_noise=0.0, score_noise=0.3, with_probs=False) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> summary = dmet.summarize(plot=True, title='DetectionMetrics summary demo', with_ovr=True, with_bin=False) >>> summary['bin_measures'] >>> kwplot.show_if_requested()
- summarize(out_dpath=None, plot=False, title='', with_bin='auto', with_ovr='auto')[source]¶
Create summary one-versus-rest and binary metrics.
- Parameters:
out_dpath (pathlib.Path | None) – FIXME: not hooked up
with_ovr (str | bool) – include one-versus-rest metrics (wrt the classes). If ‘auto’ enables if possible. FIXME: auto is not working.
with_bin (str | bool) – include binary classless metrics (i.e. detected or not). If ‘auto’ enables if possible. FIXME: auto is not working.
plot (bool) – if true, also write plots. Defaults to False.
title (str) – passed if plot is given
Example
>>> from kwcoco.metrics.confusion_vectors import * # NOQA >>> from kwcoco.metrics.detect_metrics import DetectionMetrics >>> dmet = DetectionMetrics.demo( >>> n_fp=(0, 128), n_fn=(0, 4), nimgs=512, nboxes=(0, 32), >>> classes=3, rng=0) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> dmet.summarize(plot=True, title='DetectionMetrics summary demo') >>> kwplot.show_if_requested()
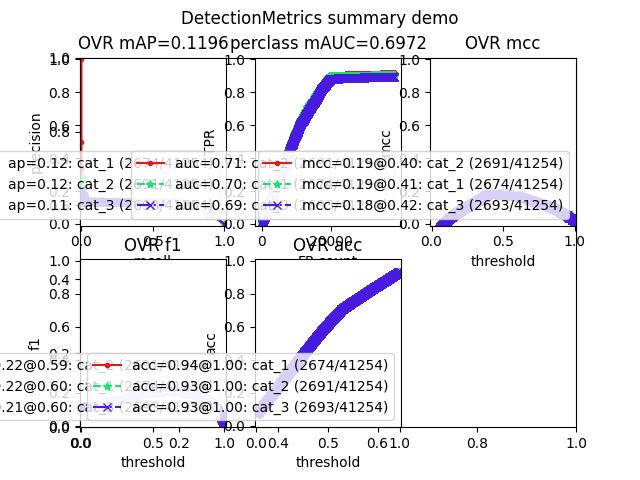
- class kwcoco.metrics.Measures(info)[source]¶
-
Holds accumulated confusion counts, and derived measures
Example
>>> from kwcoco.metrics.confusion_vectors import BinaryConfusionVectors # NOQA >>> binvecs = BinaryConfusionVectors.demo(n=100, p_error=0.5) >>> self = binvecs.measures() >>> print('self = {!r}'.format(self)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw(doclf=True) >>> self.draw(key='pr', pnum=(1, 2, 1)) >>> self.draw(key='roc', pnum=(1, 2, 2)) >>> kwplot.show_if_requested()
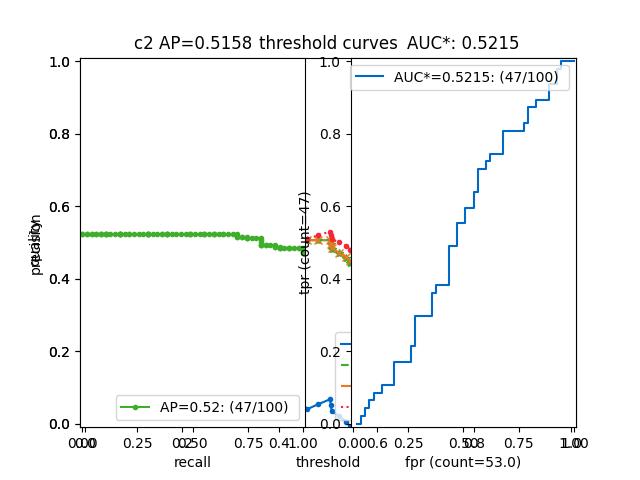
- property catname¶
- draw(key=None, prefix='', **kw)[source]¶
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(module:pandas) >>> from kwcoco.metrics.confusion_vectors import ConfusionVectors # NOQA >>> cfsn_vecs = ConfusionVectors.demo() >>> ovr_cfsn = cfsn_vecs.binarize_ovr(keyby='name') >>> self = ovr_cfsn.measures()['perclass'] >>> self.draw('mcc', doclf=True, fnum=1) >>> self.draw('pr', doclf=1, fnum=2) >>> self.draw('roc', doclf=1, fnum=3)
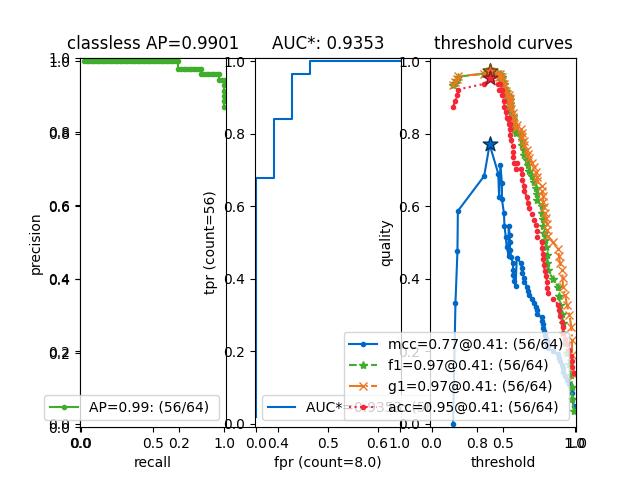
- summary_plot(fnum=1, title='', subplots='auto')[source]¶
Example
>>> from kwcoco.metrics.confusion_measures import * # NOQA >>> from kwcoco.metrics.confusion_vectors import ConfusionVectors # NOQA >>> cfsn_vecs = ConfusionVectors.demo(n=3, p_error=0.5) >>> binvecs = cfsn_vecs.binarize_classless() >>> self = binvecs.measures() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.summary_plot() >>> kwplot.show_if_requested()
- classmethod demo(**kwargs)[source]¶
Create a demo Measures object for testing / demos
- Parameters:
**kwargs – passed to
BinaryConfusionVectors.demo(). some valid keys are: n, rng, p_rue, p_error, p_miss.
- classmethod combine(tocombine, precision=None, growth=None, thresh_bins=None)[source]¶
Combine binary confusion metrics
- Parameters:
tocombine (List[Measures]) – a list of measures to combine into one
precision (int | None) – If specified rounds thresholds to this precision which can prevent a RAM explosion when combining a large number of measures. However, this is a lossy operation and will impact the underlying scores. NOTE: use
growthinstead.growth (int | None) – if specified this limits how much the resulting measures are allowed to grow by. If None, growth is unlimited. Otherwise, if growth is ‘max’, the growth is limited to the maximum length of an input. We might make this more numerical in the future.
thresh_bins (int | None) – Force this many threshold bins.
- Returns:
kwcoco.metrics.confusion_measures.Measures
Example
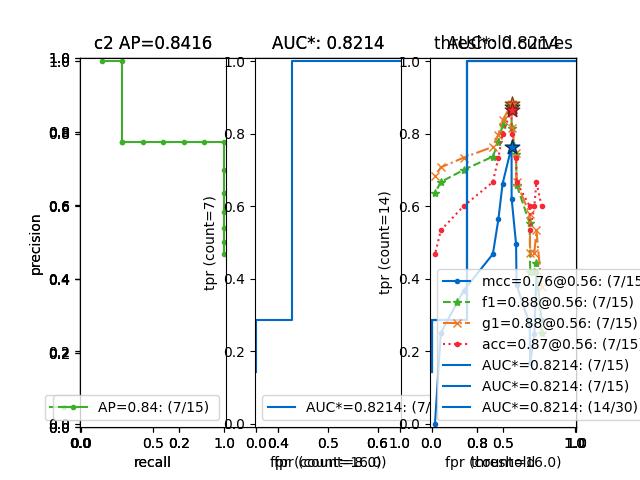
>>> from kwcoco.metrics.confusion_measures import * # NOQA >>> measures1 = Measures.demo(n=15) >>> measures2 = measures1 >>> tocombine = [measures1, measures2] >>> new_measures = Measures.combine(tocombine) >>> new_measures.reconstruct() >>> print('new_measures = {!r}'.format(new_measures)) >>> print('measures1 = {!r}'.format(measures1)) >>> print('measures2 = {!r}'.format(measures2)) >>> print(ub.urepr(measures1.__json__(), nl=1, sort=0)) >>> print(ub.urepr(measures2.__json__(), nl=1, sort=0)) >>> print(ub.urepr(new_measures.__json__(), nl=1, sort=0)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1) >>> new_measures.summary_plot() >>> measures1.summary_plot() >>> measures1.draw('roc') >>> measures2.draw('roc') >>> new_measures.draw('roc')
Example
>>> # Demonstrate issues that can arrise from choosing a precision >>> # that is too low when combining metrics. Breakpoints >>> # between different metrics can get muddled, but choosing a >>> # precision that is too high can overwhelm memory. >>> from kwcoco.metrics.confusion_measures import * # NOQA >>> base = ub.map_vals(np.asarray, { >>> 'tp_count': [ 1, 1, 2, 2, 2, 2, 3], >>> 'fp_count': [ 0, 1, 1, 2, 3, 4, 5], >>> 'fn_count': [ 1, 1, 0, 0, 0, 0, 0], >>> 'tn_count': [ 5, 4, 4, 3, 2, 1, 0], >>> 'thresholds': [.0, .0, .0, .0, .0, .0, .0], >>> }) >>> # Make tiny offsets to thresholds >>> rng = kwarray.ensure_rng(0) >>> n = len(base['thresholds']) >>> offsets = [ >>> sorted(rng.rand(n) * 10 ** -rng.randint(4, 7))[::-1] >>> for _ in range(20) >>> ] >>> tocombine = [] >>> for offset in offsets: >>> base_n = base.copy() >>> base_n['thresholds'] += offset >>> measures_n = Measures(base_n).reconstruct() >>> tocombine.append(measures_n) >>> for precision in [6, 5, 2]: >>> combo = Measures.combine(tocombine, precision=precision).reconstruct() >>> print('precision = {!r}'.format(precision)) >>> print('combo = {}'.format(ub.urepr(combo, nl=1))) >>> print('num_thresholds = {}'.format(len(combo['thresholds']))) >>> for growth in [None, 'max', 'log', 'root', 'half']: >>> combo = Measures.combine(tocombine, growth=growth).reconstruct() >>> print('growth = {!r}'.format(growth)) >>> print('combo = {}'.format(ub.urepr(combo, nl=1))) >>> print('num_thresholds = {}'.format(len(combo['thresholds']))) >>> #print(combo.counts().pandas())
Example
>>> # Test case: combining a single measures should leave it unchanged >>> from kwcoco.metrics.confusion_measures import * # NOQA >>> measures = Measures.demo(n=40, p_true=0.2, p_error=0.4, p_miss=0.6) >>> df1 = measures.counts().pandas().fillna(0) >>> print(df1) >>> tocombine = [measures] >>> combo = Measures.combine(tocombine) >>> df2 = combo.counts().pandas().fillna(0) >>> print(df2) >>> assert np.allclose(df1, df2)
>>> combo = Measures.combine(tocombine, thresh_bins=2) >>> df3 = combo.counts().pandas().fillna(0) >>> print(df3)
>>> # I am NOT sure if this is correct or not >>> thresh_bins = 20 >>> combo = Measures.combine(tocombine, thresh_bins=thresh_bins) >>> df4 = combo.counts().pandas().fillna(0) >>> print(df4)
>>> combo = Measures.combine(tocombine, thresh_bins=np.linspace(0, 1, 20)) >>> df4 = combo.counts().pandas().fillna(0) >>> print(df4)
assert np.allclose(combo[‘thresholds’], measures[‘thresholds’]) assert np.allclose(combo[‘fp_count’], measures[‘fp_count’]) assert np.allclose(combo[‘tp_count’], measures[‘tp_count’]) assert np.allclose(combo[‘tp_count’], measures[‘tp_count’])
globals().update(xdev.get_func_kwargs(Measures.combine))
Example
>>> # Test degenerate case >>> from kwcoco.metrics.confusion_measures import * # NOQA >>> tocombine = [ >>> {'fn_count': [0.0], 'fp_count': [359980.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [7747.0]}, >>> {'fn_count': [0.0], 'fp_count': [360849.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [424.0]}, >>> {'fn_count': [0.0], 'fp_count': [367003.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [991.0]}, >>> {'fn_count': [0.0], 'fp_count': [367976.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [1017.0]}, >>> {'fn_count': [0.0], 'fp_count': [676338.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [7067.0]}, >>> {'fn_count': [0.0], 'fp_count': [676348.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [7406.0]}, >>> {'fn_count': [0.0], 'fp_count': [676626.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [7858.0]}, >>> {'fn_count': [0.0], 'fp_count': [676693.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [10969.0]}, >>> {'fn_count': [0.0], 'fp_count': [677269.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [11188.0]}, >>> {'fn_count': [0.0], 'fp_count': [677331.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [11734.0]}, >>> {'fn_count': [0.0], 'fp_count': [677395.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [11556.0]}, >>> {'fn_count': [0.0], 'fp_count': [677418.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [11621.0]}, >>> {'fn_count': [0.0], 'fp_count': [677422.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [11424.0]}, >>> {'fn_count': [0.0], 'fp_count': [677648.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [9804.0]}, >>> {'fn_count': [0.0], 'fp_count': [677826.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [2470.0]}, >>> {'fn_count': [0.0], 'fp_count': [677834.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [2470.0]}, >>> {'fn_count': [0.0], 'fp_count': [677835.0], 'thresholds': [0.0], 'tn_count': [0.0], 'tp_count': [2470.0]}, >>> {'fn_count': [11123.0, 0.0], 'fp_count': [0.0, 676754.0], 'thresholds': [0.0002442002442002442, 0.0], 'tn_count': [676754.0, 0.0], 'tp_count': [2.0, 11125.0]}, >>> {'fn_count': [7738.0, 0.0], 'fp_count': [0.0, 676466.0], 'thresholds': [0.0002442002442002442, 0.0], 'tn_count': [676466.0, 0.0], 'tp_count': [0.0, 7738.0]}, >>> {'fn_count': [8653.0, 0.0], 'fp_count': [0.0, 676341.0], 'thresholds': [0.0002442002442002442, 0.0], 'tn_count': [676341.0, 0.0], 'tp_count': [0.0, 8653.0]}, >>> ] >>> thresh_bins = np.linspace(0, 1, 4) >>> combo = Measures.combine(tocombine, thresh_bins=thresh_bins).reconstruct() >>> print('tocombine = {}'.format(ub.urepr(tocombine, nl=2))) >>> print('thresh_bins = {!r}'.format(thresh_bins)) >>> print(ub.urepr(combo.__json__(), nl=1)) >>> for thresh_bins in [4096, 1]: >>> combo = Measures.combine(tocombine, thresh_bins=thresh_bins).reconstruct() >>> print('thresh_bins = {!r}'.format(thresh_bins)) >>> print('combo = {}'.format(ub.urepr(combo, nl=1))) >>> print('num_thresholds = {}'.format(len(combo['thresholds']))) >>> for precision in [6, 5, 2]: >>> combo = Measures.combine(tocombine, precision=precision).reconstruct() >>> print('precision = {!r}'.format(precision)) >>> print('combo = {}'.format(ub.urepr(combo, nl=1))) >>> print('num_thresholds = {}'.format(len(combo['thresholds']))) >>> for growth in [None, 'max', 'log', 'root', 'half']: >>> combo = Measures.combine(tocombine, growth=growth).reconstruct() >>> print('growth = {!r}'.format(growth)) >>> print('combo = {}'.format(ub.urepr(combo, nl=1))) >>> print('num_thresholds = {}'.format(len(combo['thresholds'])))
- class kwcoco.metrics.OneVsRestConfusionVectors(cx_to_binvecs, classes)[source]¶
Bases:
NiceReprContainer for multiple one-vs-rest binary confusion vectors
- Variables:
cx_to_binvecs –
classes –
Example
>>> from kwcoco.metrics import DetectionMetrics >>> dmet = DetectionMetrics.demo( >>> nimgs=10, nboxes=(0, 10), n_fp=(0, 1), classes=3) >>> cfsn_vecs = dmet.confusion_vectors() >>> self = cfsn_vecs.binarize_ovr(keyby='name') >>> print('self = {!r}'.format(self))
- classmethod demo()[source]¶
- Parameters:
**kwargs – See
kwcoco.metrics.DetectionMetrics.demo()- Returns:
ConfusionVectors
- measures(stabalize_thresh=7, fp_cutoff=None, monotonic_ppv=True, ap_method='pycocotools')[source]¶
Creates binary confusion measures for every one-versus-rest category.
- Parameters:
stabalize_thresh (int) – if fewer than this many data points inserts dummy stabilization data so curves can still be drawn. Default to 7.
fp_cutoff (int | None) – maximum number of false positives in the truncated roc curves. The default
Noneis equivalent tofloat('inf')monotonic_ppv (bool) – if True ensures that precision is always increasing as recall decreases. This is done in pycocotools scoring, but I’m not sure its a good idea. Default to True.
Example
>>> self = OneVsRestConfusionVectors.demo() >>> thresh_result = self.measures()['perclass']
- class kwcoco.metrics.PerClass_Measures(cx_to_info)[source]¶
-
- draw(key='mcc', prefix='', **kw)[source]¶
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwcoco.metrics.confusion_vectors import ConfusionVectors # NOQA >>> cfsn_vecs = ConfusionVectors.demo() >>> ovr_cfsn = cfsn_vecs.binarize_ovr(keyby='name') >>> self = ovr_cfsn.measures()['perclass'] >>> self.draw('mcc', doclf=True, fnum=1) >>> self.draw('pr', doclf=1, fnum=2) >>> self.draw('roc', doclf=1, fnum=3)
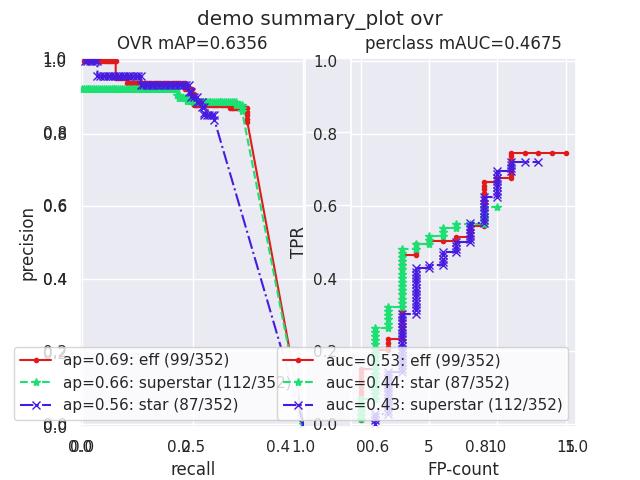
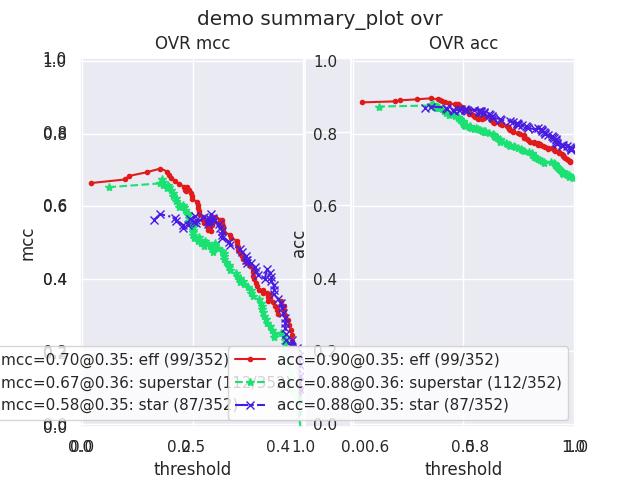
- summary_plot(fnum=1, title='', subplots='auto')[source]¶
CommandLine
python ~/code/kwcoco/kwcoco/metrics/confusion_measures.py PerClass_Measures.summary_plot --show
Example
>>> from kwcoco.metrics.confusion_measures import * # NOQA >>> from kwcoco.metrics.detect_metrics import DetectionMetrics >>> dmet = DetectionMetrics.demo( >>> n_fp=(0, 1), n_fn=(0, 3), nimgs=32, nboxes=(0, 32), >>> classes=3, rng=0, newstyle=1, box_noise=0.7, cls_noise=0.2, score_noise=0.3, with_probs=False) >>> cfsn_vecs = dmet.confusion_vectors() >>> ovr_cfsn = cfsn_vecs.binarize_ovr(keyby='name', ignore_classes=['vector', 'raster']) >>> self = ovr_cfsn.measures()['perclass'] >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> import seaborn as sns >>> sns.set() >>> self.summary_plot(title='demo summary_plot ovr', subplots=['pr', 'roc']) >>> kwplot.show_if_requested() >>> self.summary_plot(title='demo summary_plot ovr', subplots=['mcc', 'acc'], fnum=2)